Autosomal Ancestry: MODERN: --------------------------------------------------------------- My DNA Origins: --------------------------------------------------------------- My DNA Origins by GPS Origin Test Results : Overall Ancestry Origins #1 Southeastern India 42.1% Origin: Endemic to south eastern india with residues in Pakistan #2 Southwestern India 16.5% Origin: Endemic to Indian (Pulayar) with residues in India (Paniya, Savara, Bengali, Juang, Savara, Ho, Bonda) #3 Western Siberia 6.6% Origin: Peaks in Krasnoyarsk Krai and declines towards east Russia #4 Northern India 6.3% Origin: Peaks in North India (Dharkars, Kanjars) and declines in Pakistan #5 Tuva (Russia Tuva) 5.6% Origin: Peaks in south Siberia (Russians: Tuvinian) and declines in North Mongolia #6 Austronesian Southeast Asia 4.7% Origin: Peaks in Taiwan and Malay and declines in Thailand, Vietnam, Cambodia, and South China #7 Austronesian Oceania 3.3% Origin: Peaks in Korea, Chinese (Han), Mynamar, Japan, and Vietnam and declines towards West China and India #8 Bougainville 3.1% Origin: Peaks in Bougainville and declines in Australia #9 Fennoscandia 2.5% Origin: Peaks in the Iceland and Norway and declines in Finland, England, and France #10 Pima County: The Sonora 2.3% Origin: Peaks in Central-North America and declines towards Greenland and Eskimos #11 Papuan New Guinea 1.9% Origin: Peaks in Papua New Guinea and declines in Australia #12 Basque Country 1.9% Origin: Peaks in France and Spain Basque regions and declines in Spain, France, and Germany #13 Central America 1.4% Origin: Peaks in Mexico and Central America with residues in Peru #14 Southern France 1% Origin: Peaks in south France and declines in north France, England, Orkney islands, and Scandinavia #15 The Southern Levant 0.8% Origin: This gene pool is localized to Israel with residues in Syria ------------------------------------------------------------------------------- GedMatch Ancestry Calculators Modern: Most GedMatch Ancestry Calculators : Raw File: My 23andme ancestry Original raw data from YSEQ WGS ------------------------------------------------------------------------------- Dodecad K12b Oracle results: Modern Raw File: My 23andme ancestry Original raw data from YSEQ WGS Admix Results (sorted): # Population Percent 1 South_Asian 61.25 2 Gedrosia 29.74 3 Southeast Asian 4.8 4 East_Asian 0.7 5 Southwest_Asian 0.7 6 North_European 0.69 7 Sub-Saharan 0.57 8 Northwest_African 0.47 9 Atlantic_Med 0.35 10 East_African 0.33 11 Caucasus 0.2 12 Siberian 0.2 -------------------------------------------------------------------------------------------------------------------- Eurogenes K13 Oracle results: Modern K13 Oracle ref data revised 21 Nov 2013 Raw File: My 23andme ancestry Original raw data from YSEQ WGS Admix Results (sorted): # Population Percent 1 South_Asian 75.26 2 West_Asian 12.29 3 East_Asian 4.88 4 Oceanian 1.7 5 Red_Sea 1.37 6 Sub-Saharan 1.3 7 Northeast_African 1.18 8 Baltic 1.04 9 West_Med 0.71 10 American 0.28 ---------------------------------------------------------------------------------------------------- MDLP World-22 Oracle results: Modern Admix Results (sorted): # Population Percent 1 Indian 55.35 2 West-Asian 20.67 3 East-South-Asian 8.69 4 Indo-Iranian 6.11 5 Indo-Tibetan 2.11 6 Sub-Saharian 1.49 7 Melanesian 1.32 8 Austronesian 1.16 9 North-European-Mesolithic 0.75 10 Mesoamerican 0.69 11 Arctic-Amerind 0.48 12 South-America_Amerind 0.33 13 Samoyedic 0.28 14 North-Amerind 0.27 15 North Siberian 0.16 --------------------------------------------------------------------------------------------------------------- MDLP K23b Oracle results: MDLP K23b Oracle Rev 2014 Sep 16 Raw File: My 23andme ancestry Original raw data from YSEQ WGS Admix Results (sorted): # Population Percent 1 South_Indian 64.37 2 South_Central_Asian 23.08 3 South_East_Asian 2.99 4 Austronesian 1.61 5 Australoid 1.23 6 Paleo_Siberian 1.01 7 Subsaharian 0.96 8 Melano_Polynesian 0.85 9 North_African 0.74 10 Arctic 0.72 11 European_Early_Farmers 0.69 12 European_Hunters_Gatherers 0.64 13 Ancestral_Altaic 0.62 14 Tungus-Altaic 0.26 15 Archaic_African 0.19 -------------------------------------------------------------------------------------------------------- MDLP K16 Modern Oracle results: MDLP K16 2xOracle and OracleX4 Admix Results (sorted): # Population Percent 1 Indian 68.38 2 SouthEastAsian 13.93 3 Caucasian 5.46 4 Oceanic 3.14 5 Australian 2.9 6 NorthAfrican 2.15 7 Amerindian 0.93 8 Arctic 0.89 9 Subsaharian 0.73 10 Siberian 0.56 11 East African 0.48 12 Neolithic 0.44 ------------------------------------------------------------------------------------------- MDLP World Oracle results: Admix Results (sorted): # Population Percent 1 Indian 60.56 2 Caucaus_Parsia 23.73 3 East_Asian 7.03 4 Melanesian 2.58 5 North_and_East_European 2.51 6 Sub_Saharian 1.61 7 Mesoamerican 0.73 8 Middle_East 0.67 9 Arctic_Amerind 0.58 ------------------------------------------------------------------------------- Gedrosia K12 Oracle results: Admix Results (sorted): # Population Percent 1 S_INDIAN 66.35 2 Baloch.i 16.06 3 INDO_TIBETAN 6.16 4 CAUCASUS 3.07 5 SE_ASIAN 2.97 6 E_SIBERIAN 1.4 7 EARLY_EUROPEAN_FARMERS 1.17 8 SINTASHTA_STEPPE_HERDERS 1.13 9 E_AFRICAN 1.06 10 SUB_SAHARAN 0.63 --------------------------------------------------------------------------------------------------------------- HarappaWorld Oracle results: 23 April 2013 - Oracle reference population percentages revised. Raw File: My 23andme From YSEQ WGS raw data File further imputed Raw file from DNAGenics Admix Results (sorted): # Population Percent 1 S-Indian 60.35 2 Baloch 29.08 3 SE-Asian 4.01 4 Papuans 1.64 5 SW-Asian 1.2 6 Mediterranean 0.82 7 Beringian 0.74 8 American 0.52 9 W-African 0.35 10 Caucasian 0.19 11 NE-Euro 0.19 12 NE-Asian 0.19 13 Siberian 0.19 14 Pygmy 0.19 15 San 0.19 16 E-African 0.19 ----------------------------------------------------------------------------------------------- ANCIENT: ----------------------------------------------------------------------------------------------- GedMatch Ancestry Calculators Ancient: ----------------------------------------------------------------------------------------------- MDLP K11 Modern Oracle results: Archaic Roots MDLP K11 2xOracle Raw File: My 23andme ancestry Original raw data from YSEQ WGS Admix Results (sorted): File: 23andme # Population Percent 1 ASI 67.31 2 EHG 20.07 3 SEA 7.59 4 Oceanic 1.41 5 Basal 1.11 6 Iran-Mesolithic 0.78 7 African 0.75 8 American 0.64 9 WHG 0.33 ---------------------------------------------------------------------------------- puntDNAL K12 Ancient Oracle results: puntDNAL K12 Ancient Oracle Admix Results (sorted): # Population Percent 1 South_Asian 63.19 2 Caucasus_HG 20.55 3 East_Asian 8.95 4 Oceanian 2.94 5 European_HG 2.52 6 Sub-Saharan 0.91 7 Beringian 0.41 8 South_African_HG 0.32 9 Anatolian_NF 0.21 ------------------------------------------------------------------------- puntDNAL K10 Ancient Oracle results: puntDNAL K10 Ancient Oracle Admix Results (sorted): # Population Percent 1 ASI 62.83 2 CHG 19.77 3 E_Asian 7.84 4 Oceanian 3.1 5 Sub-Saharan 2.8 6 Beringian 1.99 7 WHG 1.56 8 ENF 0.12 ------------------------------------------------------------------------------- 2) G25 Coordinates: Ancient Ancestry ----------------------------------------------------------------------------------- a) DNAGenics Ancient Ancestry of My Overall and both Parents by Raw File Phasing of my File My Overall Ancestry file: YSEQ 23andme ancestry file G25 Co-Ordinates from DNAgenics 🧬DNAGENICS G25 Studio Results Calculator: OG Old World (Improved, Simplified) Fit Score: 0.80 Chebyshev Ancestry Composition: AASI: 56.60% Iranian Mesolithic: 11.80% Ancient Near East: 8.00% Southeast Asian Lineage: 7.20% Caucasian Hunter Gatherers: 5.40% Eastern European Hunter Gatherers: 2.60% Ancient Paleo Siberians: 2.40% Ancient Northern Eurasian: 2.20% Anatolian Farmers: 1.20% Eastern Africa Pastoral: 0.80% Australoid: 0.40% Eastern Africa: 0.40% Southern Africa: 0.40% Western European Hunter Gatherers: 0.40% Ancient Northeast Asians: 0.20% Father(Parent1): Scaled G25 Coordinates from my Raw Phasing File 🧬DNAGENICS G25 Studio Results Calculator: OG Old World (Improved, Simplified) Fit Score: 0.82 Chebyshev Ancestry Composition: AASI: 52.00% Iranian Mesolithic: 11.20% Southeast Asian Lineage: 9.80% Caucasian Hunter Gatherers: 5.80% Ancient Near East: 5.60% Anatolian Farmers: 3.00% Ancient Northern Eurasian: 3.00% Ancient Paleo Siberians: 2.60% Ancient Northeast Asians: 2.00% Eastern European Hunter Gatherers: 1.40% Western European Hunter Gatherers: 1.00% Australoid: 0.80% Eastern Africa Pastoral: 0.80% Eastern Africa: 0.40% Neolithic Yellow River: 0.40% Southern Africa: 0.20% Mother(Parent2): Scaled G25 Coordinates from my Raw Phasing File 🧬DNAGENICS G25 Studio Results Calculator: OG Old World (Improved, Simplified) Fit Score: 0.87 Chebyshev Ancestry Composition: AASI: 53.00% Iranian Mesolithic: 17.00% Ancient Near East: 5.60% Caucasian Hunter Gatherers: 4.20% Eastern European Hunter Gatherers: 4.20% Ancient Northern Eurasian: 4.00% Anatolian Farmers: 3.60% Neolithic Yellow River: 2.20% Australoid: 1.80% Southeast Asian Lineage: 1.60% Ancient Paleo Siberians: 1.20% Ancient Northeast Asians: 0.60% Southern Africa: 0.60% Proto-Bantu Africa: 0.20% Eastern Africa Pastoral: 0.20% ---------------------------------------------------------------------------------------------------------------- My Overall Ancestry file: YSEQ 23andme ancestry file G25 Co-Ordinates from DNAgenics 🧬DNAGENICS G25 Studio Results Calculator: IllustrativeDNA-Like Ancient calculator Fit Score: 0.50 Chebyshev Ancestry Composition: Indian Subcontinent (AD 690–990): 68.40% Ancient Ancestral South Indian: 15.40% Swat Valley (300 BC–AD 1350): 8.20% Southeast Asian (2000 BC–AD 1800): 2.20% Gandhara Grave Culture (1300–800 BC): 2.00% Australian (2000 BC–AD 1600): 1.20% Lazica: 1.20% Cushitic (2000 BC–AD 600): 0.40% Iranian Plateau: 0.20% Kartvelian: 0.20% Khwarazm And Transoxiana (100 BC–AD 950): 0.20% North Caucasian (AD 650–1160): 0.20% Tibetan Plateau (1200 BC–AD 500): 0.20% Father(Parent1): Scaled G25 Coordinates from my Raw Phasing File 🧬DNAGENICS G25 Studio Results Calculator: IllustrativeDNA-Like Ancient calculator Fit Score: 0.53 Chebyshev Ancestry Composition: Indian Subcontinent (AD 690–990): 66.40% Ancient Ancestral South Indian: 15.20% Swat Valley (300 BC–AD 1350): 10.20% Southeast Asian (2000 BC–AD 1800): 4.80% Australian (2000 BC–AD 1600): 1.20% Gandhara Grave Culture (1300–800) BC): 1.20% Kartvelian: 0.20% Lazica: 0.20% Old Bering Sea Culture (AD 200–1330): 0.20% Tibetan Plateau (1200 BC–AD 500) : 0.20% Turkic (AD 650–1200) : 0.20% Mother(Parent2): Scaled G25 Coordinates from my Raw Phasing File 🧬DNAGENICS G25 Studio Results Calculator: IllustrativeDNA-Like Ancient calculator Fit Score: 0.54 Chebyshev Ancestry Composition: Indian Subcontinent (AD 690–990): 51.80% Swat Valley (300 BC–AD 1350): 21.20% Ancient Ancestral South Indian: 16.80% Gandhara Grave Culture (1300–800 BC): 8.40% Australian (2000 BC–AD 1600): 0.80% Southeast Asian (2000 BC–AD 1800): 0.80% Lazica: 0.20% ----------------------------------------------------------------------------------------------- Paternal Side Gedmatch matches : ALL IBD GedMatch Top 5 Varied Match Results Compared to my Indian Ancestry with one being Focal Tamilian Connection and others possible Buddhist and Non-Buddhist Connections: Ethnicity Based on Eurogenes K13 results and centimorgan(cM) Match "Identical by Descant" by Q Score 1) Xibe-Uyghur + Anglo-Burmese : Total cM -10.5 cm, Largest cM 10.5 cm - Male Most Recent Common Ancestor (MRCA) - Gen 5.21 "Identical by Descant" by Q Score Plus Confirmed Relative via GedMatch subscribed tool Auto-Kinship. My GedMatch Match Rank : 8 Possible Buddhist Connection from one or two lines of this mixed ancestral relative match. Focal) Tamilian: Total cM -29.7 cM, Largest cM 18.3 cM - Male Most Recent Common Ancestor (MRCA) - Gen 4.46 "Identical by Descant" by Q Score My GedMatch Match Rank : 1 Focal Connection : Strongly Connected by Paternal Indian caste "Tamil Sakilli" 2) Manchu Chinese: Total cM -12.3 cM, Largest cM 12.3 cM - Male Most Recent Common Ancestor (MRCA) - Gen 5.10 "Identical by Descant" by Q Score My GedMatch Match Rank : 2 Possible Buddhist Connection 3) Han Chinese: Total cM -11.5 cM, Largest cM 11.5cM - Female Most Recent Common Ancestor (MRCA) - Gen 5.14 "Identical by Descant" by Q Score My GedMatch Match Rank : 4 Possible Buddhist Connection 4) South West English: Total cM -11.6 cM, Largest cM 11.6 cM - Female Most Recent Common Ancestor (MRCA) - Gen 5.14 "Identical by Descant" by Q Score My GedMatch Match Rank : 3 Non Buddhist Connection 5) Uyghurs Profile(Uyghur-Hui??) : Total cM -10.7 cM, Largest cM 10.7 cM - Female Most Recent Common Ancestor (MRCA) - Gen 5.20 "Identical by Descant" by Q Score My GedMatch Match Rank : 7 Non-Buddhist Connection My Paternal Side Ancestry relevant Population Distance including Proxy Populations and Admixture Components by each Gedmatch Calculator predictions: (In Progress) Note: Proxy Populations are marked as Proxy in Brackets. Relevant Paternal Genetically direct applicable Populations are unmarked 1) HarappaWorld Single Population Sharing: Population (source) Distance tamil-nadu-scheduled-caste (metspa.lu) 4.77 tharu (rei.ch) 7.69 Sakilli (Chaubey) 7.81 Mixed Mode Population Sharing Primary Population (source) Secondary Population (source) Distance 96.9% singapore-indian-a (sgvp)(Proxy) + 3.1% iban (xing) @ 1.98 96.1% singapore-indian-a (sgvp)(Proxy) + 3.9% thai (xing) @ 2.21 2)MDLP K16 Modern Single Population Sharing: Population (source) Distance Sakilli (Tamil_Nadu) 3.07 Scheduled_Caste (Tamil_Nadu) 6.59 3)MDLP World-22 Single Population Sharing: Population (source) Distance Hindu (derived) 8.02 Indian (derived) 9.09 Indian (ancestral) 44.59 Tadjik (derived) 46.11 Hazara (derived) 46.82 Uzbek (derived) 47.65 Turkmen (derived) 47.97 Uyghur (derived) 50.8 Iranian (derived) 54.05 Karakalpak (derived) 55.23 Mixed Mode Population Sharing Primary Population (source) Secondary Population (source) Distance 93.7% Hindu (derived) + 6.3% Dai (derived) @ 4.84 93.7% Hindu (derived) + 6.3% Lahu (derived) @ 4.86 93.6% Hindu (derived) + 6.4% She (derived) @ 5 93.5% Hindu (derived) + 6.5% Chinese-South (derived) @ 5.1 91.4% Hindu (derived) + 8.6% Burma (derived) @ 5.02 93.4% Hindu (derived) + 6.6% Han (derived) @ 5.25 93.4% Hindu (derived) + 6.6% Han-Beijing (derived) @ 5.38 93.3% Hindu (derived) + 6.7% Naxi (derived) @ 5.45 92.2% Indian (derived) + 7.8% West-Asian (ancestral) @ 5.57 4)MDLP K23b Single Population Sharing: Population (source) Distance Scheduled_Caste_Tamil_Nadu ( ) 4.91 Tharu ( ) 6.07 Mixed Mode Population Sharing Primary Population (source) Secondary Population (source) Distance 95.1% Scheduled_Caste_Tamil_Nadu ( ) + 4.9% Tai_Yuan ( ) @ 2.14 95.3% Scheduled_Caste_Tamil_Nadu ( ) + 4.7% Cantonese ( ) @ 2.16 95.1% Scheduled_Caste_Tamil_Nadu ( ) + 4.9% Wa ( ) @ 2.16 95.2% Scheduled_Caste_Tamil_Nadu ( ) + 4.8% Han_Singapore ( ) @ 2.16 95.3% Scheduled_Caste_Tamil_Nadu ( ) + 4.7% Tai_Khuen ( ) @ 2.17 94.8% Scheduled_Caste_Tamil_Nadu ( ) + 5.2% Karen ( ) @ 2.18 95.2% Scheduled_Caste_Tamil_Nadu ( ) + 4.8% Chinese_Taiwan ( ) @ 2.18 95.3% Scheduled_Caste_Tamil_Nadu ( ) + 4.7% Thai_Lue ( ) @ 2.18 95.4% Scheduled_Caste_Tamil_Nadu ( ) + 4.6% She ( ) @ 2.19 95.4% Scheduled_Caste_Tamil_Nadu ( ) + 4.6% Han ( ) @ 2.19 95.3% Scheduled_Caste_Tamil_Nadu ( ) + 4.7% Hakka ( ) @ 2.19 5)MDLP World Single Population Sharing: Population (source) Distance Indian 5.47 Tajik 49.62 Kusunda(Nepali)49.71 Hazara 51.7 Turkmen 51.94 Uzbek 52.19 Uyghur 56.51 6)Eurogenes K13 Single Population Sharing: Population (source) Distance Sakilli 2.62 Mixed Mode Population Sharing Primary Population (source) Secondary Population (source) Distance 98.2% Sakilli + 1.8% Dai @ 1.7 98.2% Sakilli + 1.8% She @ 1.72 98.1% Sakilli + 1.9% Lahu @ 1.72 97.5% Sakilli + 2.5% Tibeto-Burman_Burmese @ 1.74 98.1% Sakilli + 1.9% Naxi @ 1.81 98% Sakilli + 2% Tu @ 1.85 98.1% Sakilli + 1.9% Japanese @ 1.86 98.2% Sakilli + 1.8% Xibo @ 1.96 98.2% Sakilli + 1.8% Hezhen @ 1.97 97.5% Sakilli + 2.5% Uyghur @ 2.01 7)Dodecad K12b Single Population Sharing: Population (source) Distance Tamil_Nadu_Scheduled_Caste (Metspa.lu) 5.31 Tharus (Metspa.lu) 7.97 SAKILLI (Need) 10.8 Mixed Mode Population Sharing Primary Population (source) Secondary Population (source) Distance 93.2% Piramalai_Kallars (Metspalu) (Proxy) + 6.8% MAS30 (SGVP) @ 2.28 94.5% Kurumba (Metspalu) (Proxy) + 5.5% Dai (HGDP) @ 2.29 93.7% Piramalai_Kallars (Metspalu) (Proxy) + 6.3% CDX30 (1000Genomes) @ 2.29 94.4% Kurumba (Metspalu) (Proxy) + 5.6% CDX30 (1000Genomes) @ 2.34 8) Gedrosia K12 Single Population Sharing: Population (source) Distance Tharu 8.42 GujaratiD 8.85 Nepali 26.72 Mixed Mode Population Sharing Primary Population (source) Secondary Population (source) Distance 92.8% Velamas (Proxy) + 7.2% Kusunda (Nepali) @ 3.74 93.6% GujaratiD (Proxy) + 6.4% Push (Nepali) @ 5.22 92.2% Velamas (Proxy) + 7.8% Naga (Sino-Tibetan) @ 91.2% Kurumba (Proxy) + 8.8% Kusunda(Nepali) @ 5.35 My Paternal side Known Communities: Tamilians, Himalayan Shakyas and Burmese My Paternal side actual DNA results: Tamilians(Accurately present), Himalayan Shakyas(Accurately present), Xibe-Uyghurs(Additional Component found) ,Burmese(Present As Mixed with European & Uyghur Chinese) Note : Historical Context 1)Tharus from southern Nepal and northern India, Consider themselves as Shakya descendants and are also marked by East Eurasian and Caucasian YDNA like O-M117, O-M95,C1b1a1-M356 , Q-M242,J2a-M410(xM68, M47, M67, M158) and East Asian MTDNA along with with South Asian,& Indo-Iranian Y and MTDNA Haplogroups. Possible migration to South India during British Imperial period. They are both Hindus and Buddhist 2)Tamil Sakilli belong to Tamil Nadu Caste of Leather works 3) Manchu/Uyghur/South Chinese Prescence attributed to Qing Army POW , brought by British to Nilgiris India, after Opiums wars in China in 1800's . They were released later, employed as Tea Garden & Plantation laborers and also some were in Dairy Business. Buddhism was present in Qing dynasty. 4) 1800's were marked by Anglo-Burmese war, and many displacement and migrations. Burmese were influenced by British politics from early 1800's and and finally fully occupied it in 1875. My Great Grandfather was born in Burma, but his own Grandfather(My Great Great Great Grandfather) was Buddhist and got married in India Nilgiris/Dharmapuri during Social reform period. Oral History my Buddhist ancestor was Shakya from Himalayas and fought British While majority Tamil Sakilli from Dharmapuri/Nilgiri origin is confirmed for my Dad, the one portion of Buddhist ancestry can either be of Burmese/Tai Kadai or Xibe-Uyghur-Chinese or Tahrus(Shakyas) from Himalayas. This one Buddhist ancestral origin is open case and can be one of three Tahrus(Shakyas) or Qing China POW related or Burmese or two of three mixed. ---------------------------------------- My Maternal Side Ancestry relevant Population Distance including Proxy Populations and Admixture Components by each Gedmatch Calculator predictions: (In Progress) Note: Proxy Populations are marked as Proxy in Brackets. Relevant Maternal Genetically direct applicable Populations are unmarked My Maternal side Known communities: North Karnataka Marathi & Kannada My Maternal side actual results : North Karnataka Marathi & Kannada (Calculator Proxy GIH30 + ASUR) (Proxy Accurate) --------------------------------------------------------------------------------------- Phenotypes: -------------------------------------------------------------------------------------- My Overall Ancestry file: YSEQ 23andme ancestry file G25 Co-Ordinates from DNAgenics 🧬DNAGENICS G25 Studio Results Calculator: Phenotypes Fit Score: 0.73 Ancestry Composition: Weddoi.d: 66.60% Paleo Mongoloid: 8.60% Gracile Indid: 4.60% Negrit.id:: 3.60% Armenid: 3.40% Indo Brachid: 3.20% North Pontid: 3.20% Australoid: 2.80% Orientalid: 1.40% Mtebid: 1.20% Nilotid: 0.60% Ethiopid: 0.20% Iran-Afghanistan: 0.20% Silvid: 0.20% South Sinid: 0.20% My Father (Parent 1) : DNAGenics Phasing File G25 Co-Ordinates 🧬DNAGENICS G25 Studio Results Calculator: Phenotypes Fit Score: 0.68 Ancestry Composition: Weddoi.d: 66.80% Paleo Mongoloid: 9.60% Gracile Indid: 5.60% Indo Brachid: 3.40% North Pontid: 2.80% Orientalid: 2.60% Australoid: 2.40% Mtebid: 2.40% Negrit.id 2.40% Armenid: 1.40% Ethiopid: 0.20% Melanesians: 0.20% Saharid: 0.20% My Mother (Parent 2) : DNAGenics Phasing File G25 Co-Ordinates: 🧬DNAGENICS G25 Studio Results Calculator: Phenotypes Fit Score: 0.58 Ancestry Composition: Weddoi.d: 70.00% Indo Brachid: 12.80% Nilotid: 4.00% Gracile Indid: 2.80% Paleo Mongoloid: 1.80% Australoid: 1.60% Armenid: 1.40% North Pontid: 1.40% Negrit.id: 1.20% North India: 1.00% Amazons: 0.60% Iran-Afghanistan: 0.40% Assyroid: 0.20% Bambutid: 0.20% Dinarid: 0.20% Ethiopid: 0.20% Gracile Med: 0.20% ------------------------------------------------------------------------------------------------------------------- My Overall Ancestry file: YSEQ 23andme imputed ancestry file K15-sim_unscaled G25 Coordinates 🧬DNAGENICS G25 Studio Results Calculator: Phenotypes Fit Score: 0.53 Ancestry Composition: Weddo.id: 58.60% Gracile Indid: 27.20% Indo Brachid: 4.00% Paleo Mongoloid: 3.40% Orientalid: 2.00% Mtebid: 1.20% Armenid: 0.60% Melanesians: 0.60% Negrit.id: 0.60% Australoid: 0.40% Bantuid.: 0.20% Iran-Afghanistan: 0.20% Neo Danubian: 0.20% North India: 0.20% North Pontid: 0.20% South Sinid: 0.20% Turanid: 0.20% My Father (Parent 1) Phasing-K15-sim_unscaled G25 Coordinates 🧬DNAGENICS G25 Studio Results Calculator: Phenotypes Fit Score: 0.73 Ancestry Composition: Weddo.id: 71.20% Paleo Mongoloid: 7.00% Indo Brachid: 6.20% Gracile Indid: 4.20% North Pontid: 3.00% Nilotid: 2.80% Australoid: 2.00% Negrit.id: 1.40% Amazons: 0.40% Armenid: 0.40% Dinarid: 0.40% Gracile Med: 0.40% Ethiopid: 0.20% Mtebid: 0.20% North India: 0.20% My Mother (Parent 2) Phasing-K15-sim_unscaled G25 Coordinates 🧬DNAGENICS G25 Studio Results Calculator: Phenotypes Fit Score: 0.49 Ancestry Composition: Weddo.id: 66.80% Gracile Indid: 19.20% Indo Brachid: 7.60% Paleo Mongoloid: 1.40% Nilotid: 0.80% Armenid: 0.60% North India: 0.60% North Pontid: 0.60% Aegean Med: 0.40% Australoid: 0.40% Negrit.id: 0.40% Dinarid: 0.20% Ethiopid: 0.20% Gracile Med: 0.20% Iran-Afghanistan: 0.20% Points: 0.20% South Sinid: 0.20%
YSEQ
My Y Lineage Results
My Ancient Y Lineage Origins:
YSNP Geneology:
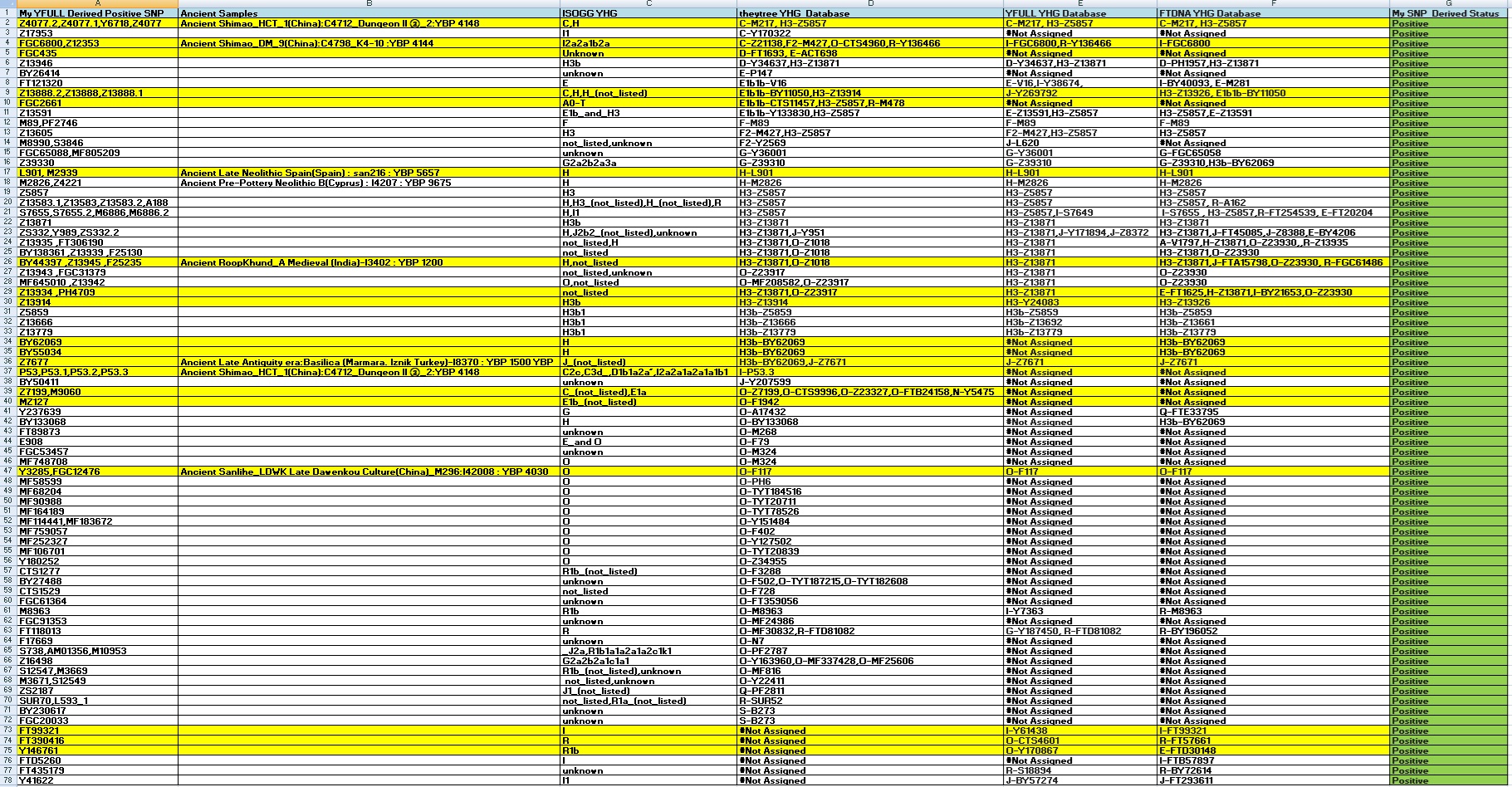
1) My YFULL BAM file Positive YSNP and its associated YHG in the theytree, YFULL and FTDNA master database and also SNP found positive in other shared YHG in research db or as private snp: :
Ancient Central Asian,SE Asian,East Asian, Caucasian,Mediterranean origins: C-M217/H3b1-BY62069/I*/J*/J-Z7671/O*/O-F117
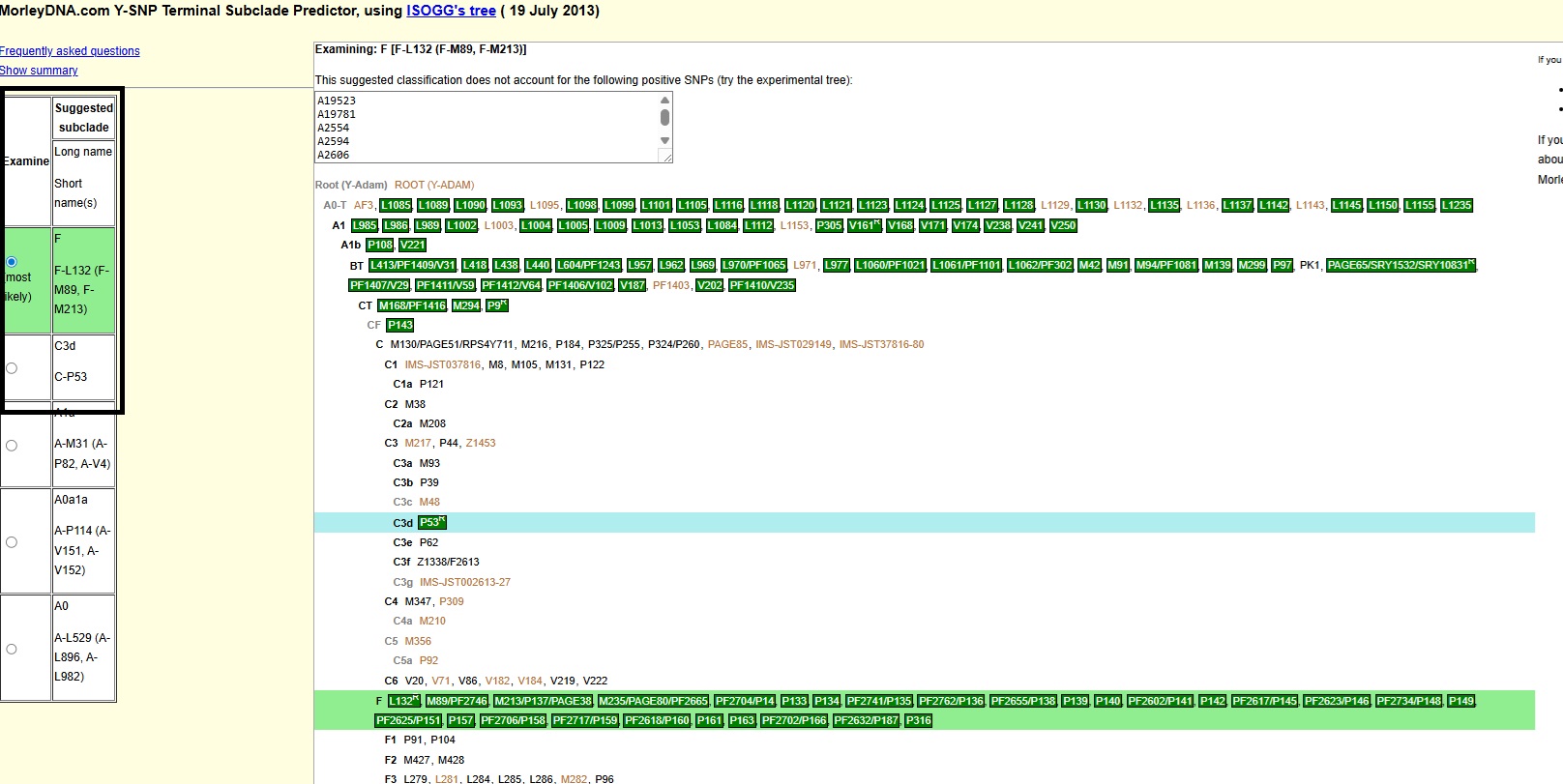
2)MorleyDNA.com Y-SNP Terminal Subclade Predictor, using ISOGG's tree (19 July 2013):
Ancient Central Asian Origins : F-M89*/C-P53
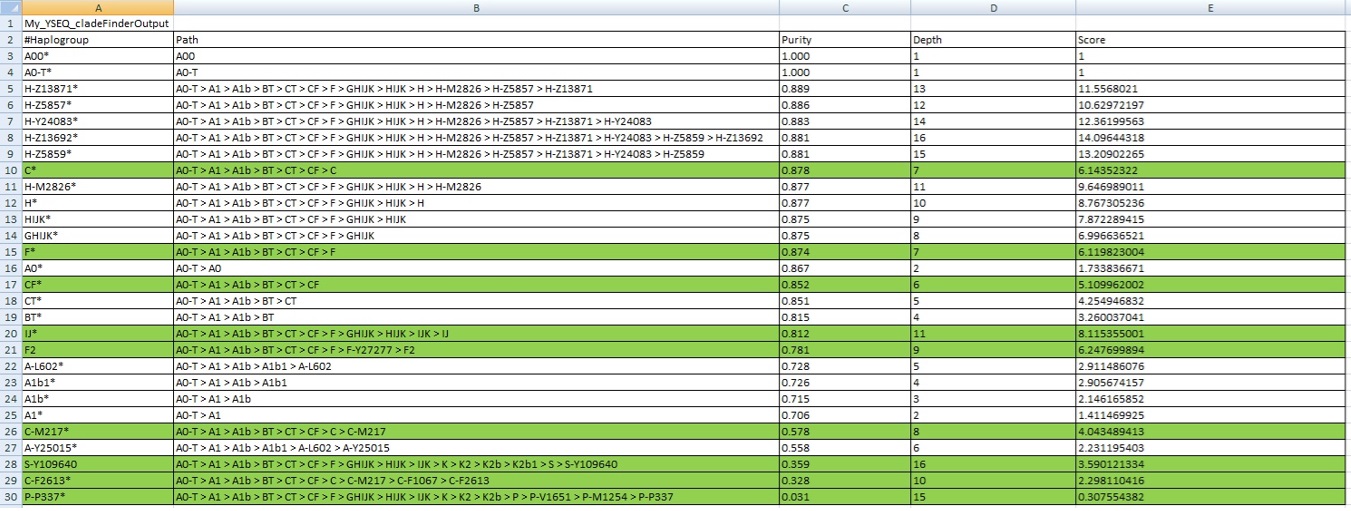
3) My YSEQ Cladefinder Output:
Ancient Caucasian,Central Asia, East Asian or Siberian Origins : YHG H3b*/C*/IJ*/F2*/C2-M217*/P1*
-----------------------------------------------------------------------
YSTR Geneology: YSEQ/YFULL/27 PCR Kit Markers
DYS393=14, DYS390=21, DYS19=14, DYS391=10
DYS385=13-16, DYS426=11, DYS388=15, DYS439=11, DYS389I=13, DYS392=11, DYS389II=30
DYS458=17,DYS459=8-8,DYS455=11,DYS454=11,DYS447=25,DYS437=14,DYS448=19,DYS449=28,
DYS464=14-15-16-17, DYS460=12,
Y-GATA-H4=11,DYS456=17,DYS576=16,DYS570=20,DYS442=12,DYS438=10,DYS531=12,DYS578=8,
DYF395=15-15,DYS590=8,DYS537=11,DYS641=10,DYS472=8,DYF406S1=12,DYS511=10,
DYS425=12,DYS413=23-23,DYS557=14,DYS436=12,DYS450=8,DYS444=12,DYS481=22,
DYS520=20,DYS617=11,DYS487=14,DYS572=11,DYS640=12,DYS492=12,
DYS565=11, DYS485=14, DYS632=8, DYS540=11, DYS556=9, DYS549=12, DYS589=14
DYS522=12, DYS494=9,
DYS533=13, DYS636=11, DYS575=10, DYS638=11, DYS462=12, DYS445=11
Y-GGAAT-1B07=10,DYS525=10,DYS650=17,DYS513=12,DYS561=14,DYS552=26,
DYS635=22, DYS587=19, DYS643=14
DYS497=15, DYS461=13, DYS435=10
My Basal Origin Minimal YSTR YHG Haplotype Prediction: Caucasian Haplotype : 12 YSTR
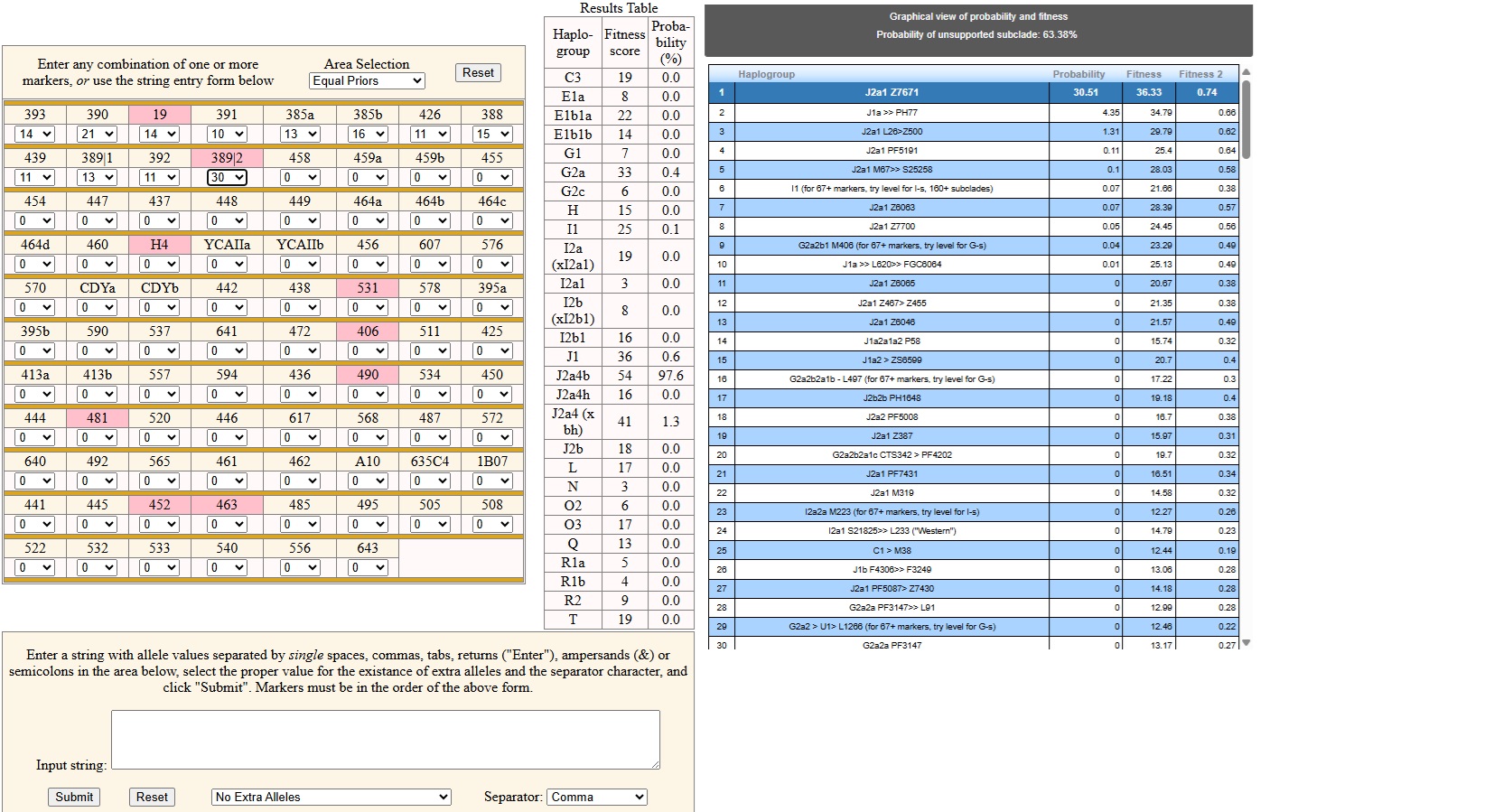
My Basal Origin First 25 YSTR YHG Haplotype Prediction: Caucasian Haplotype : 25 YSTRS
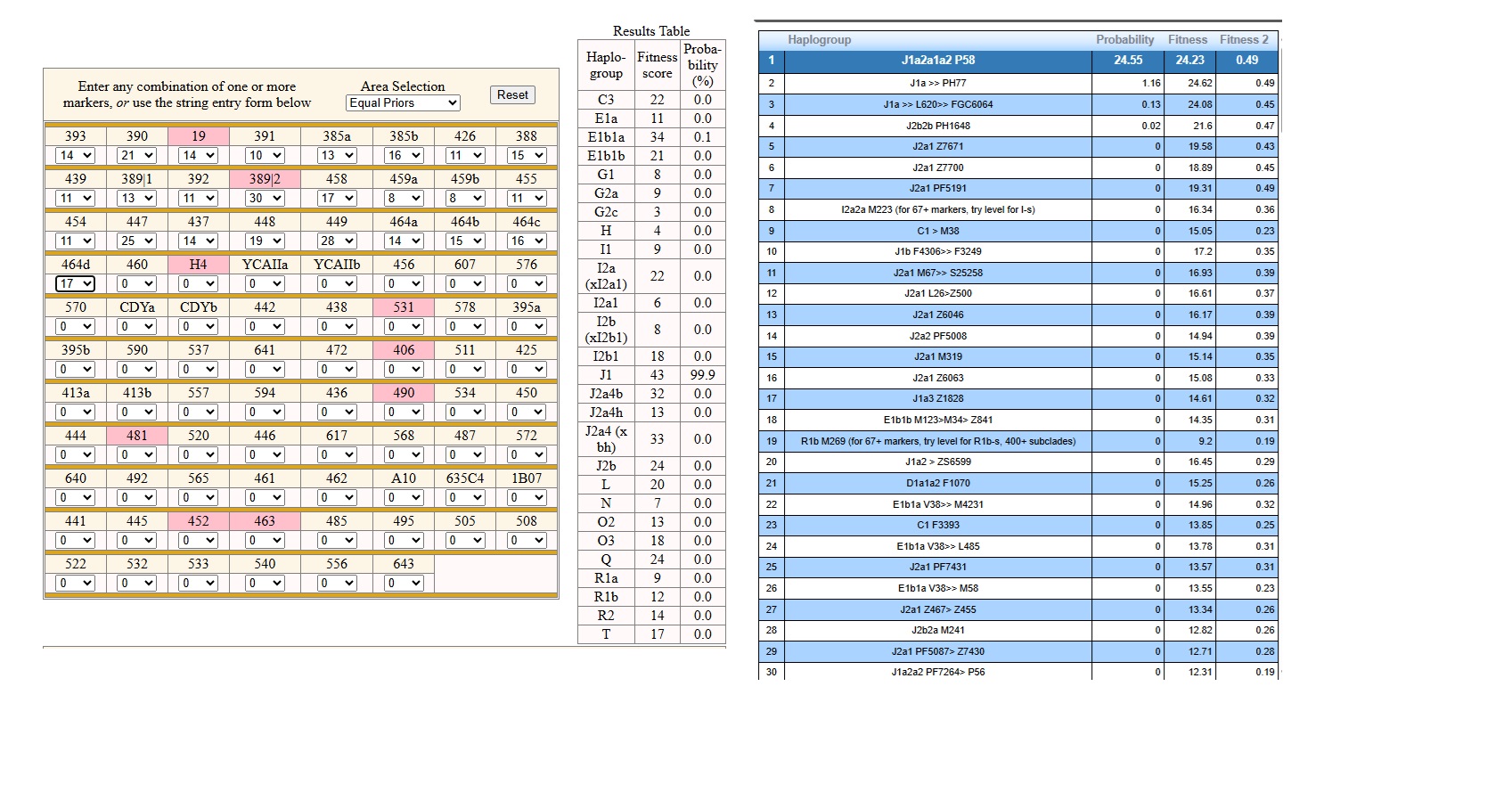
YFULL MY Sample YSTR Investigation: Analysis still in progress. Results so far with YFULL confirmed markers
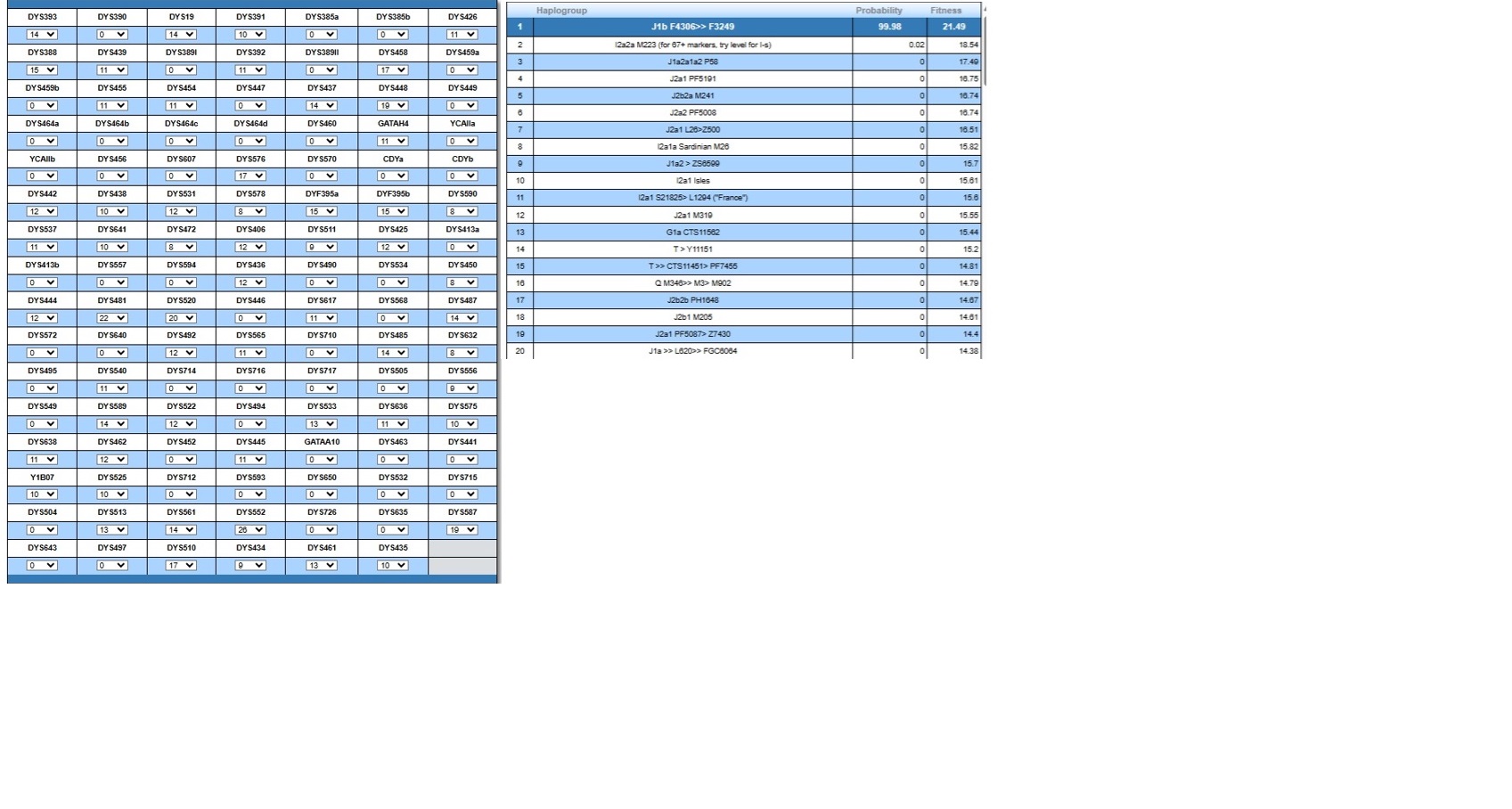
Ancient Caucasian Origin:
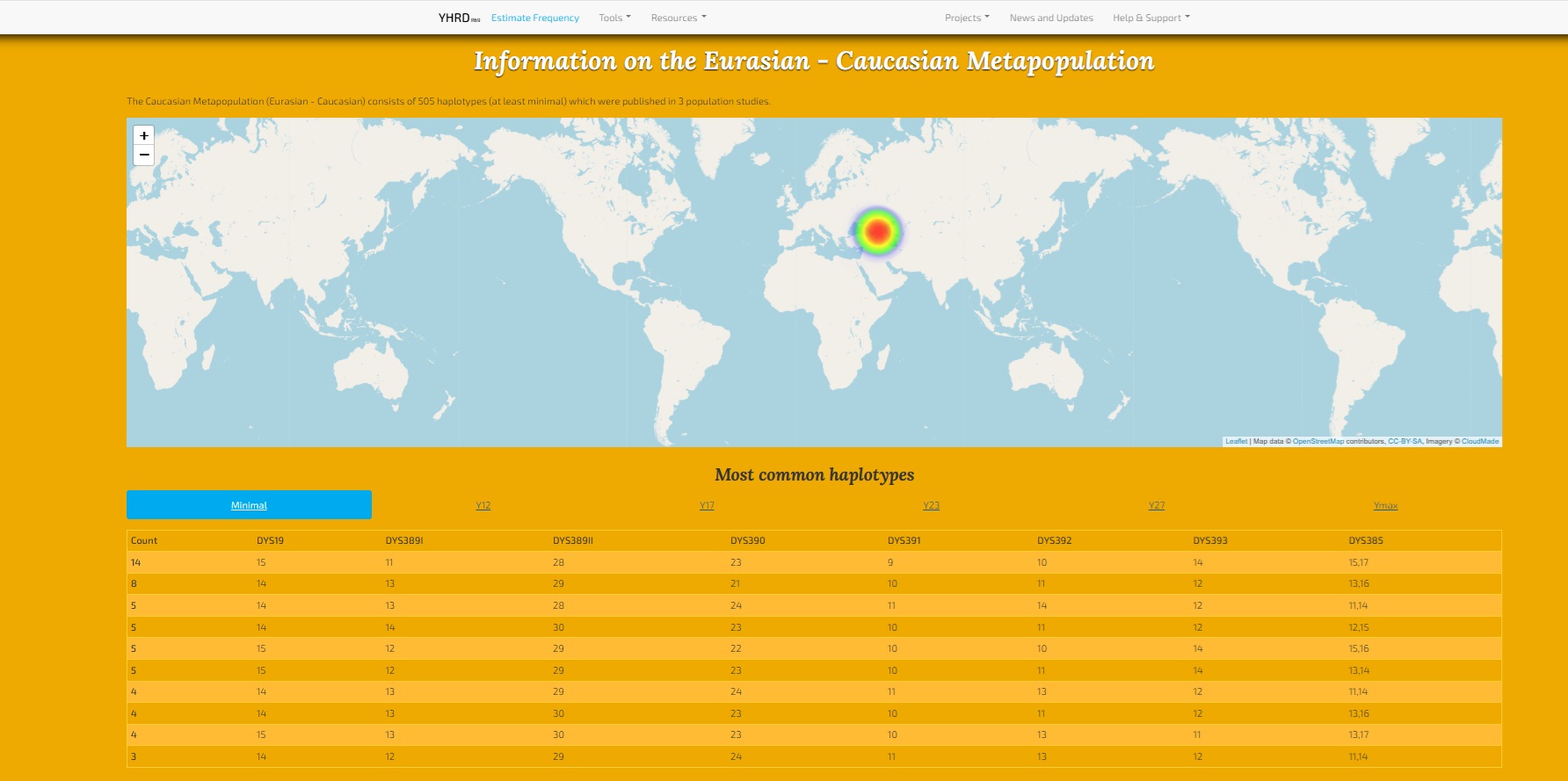
MY Sample YSTR Comparison and Matches with Close Scientific Samples:
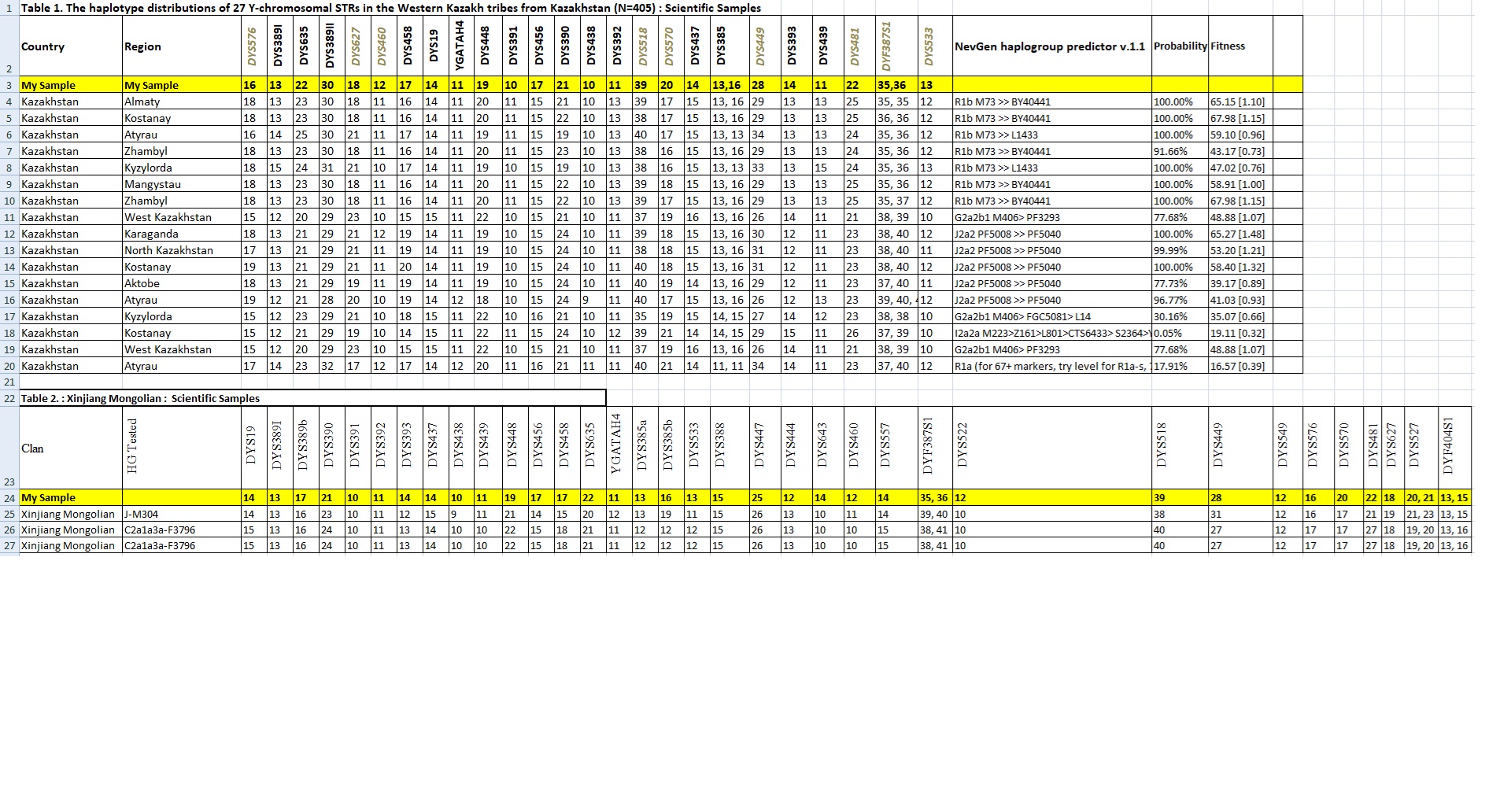
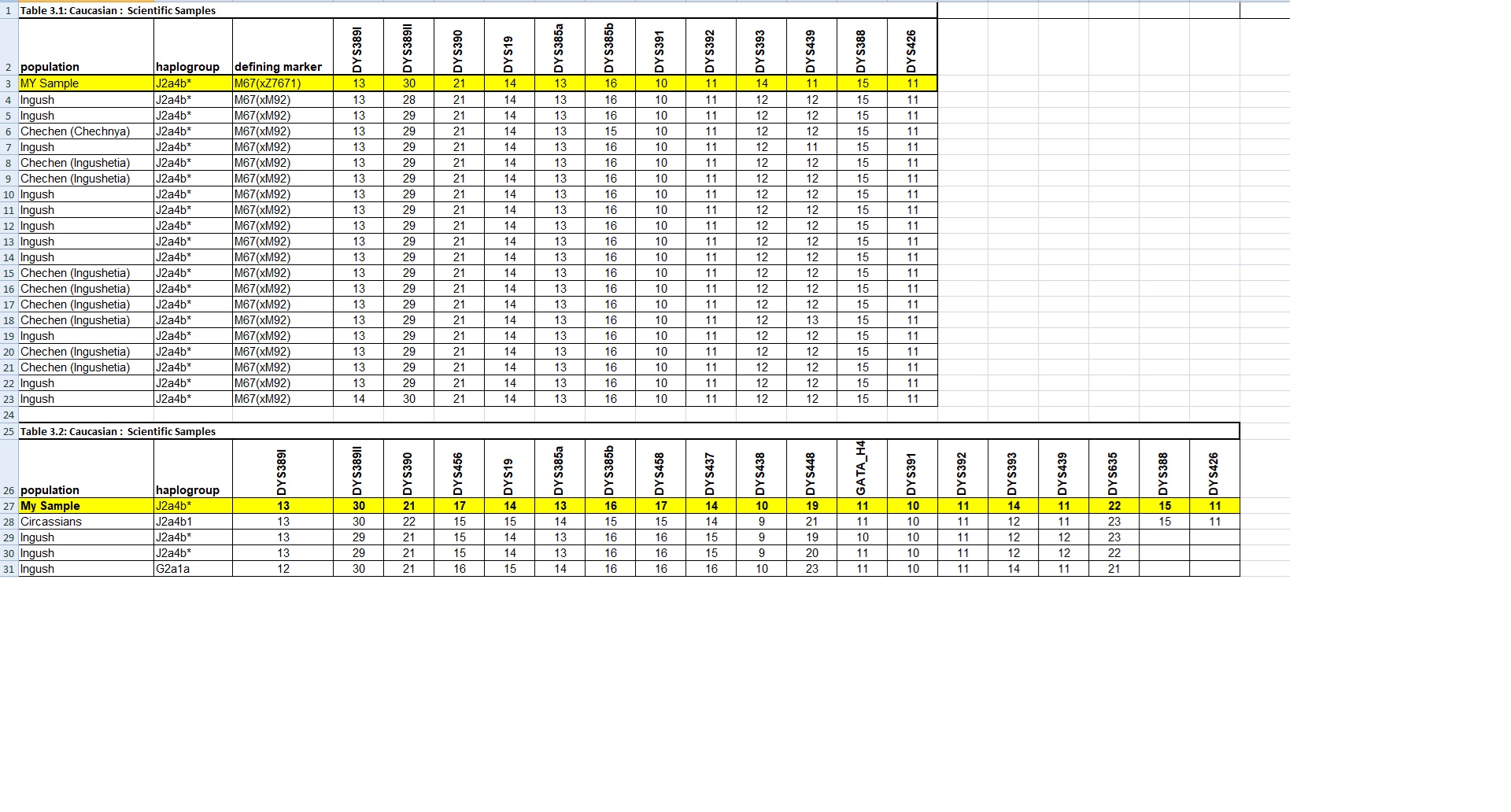
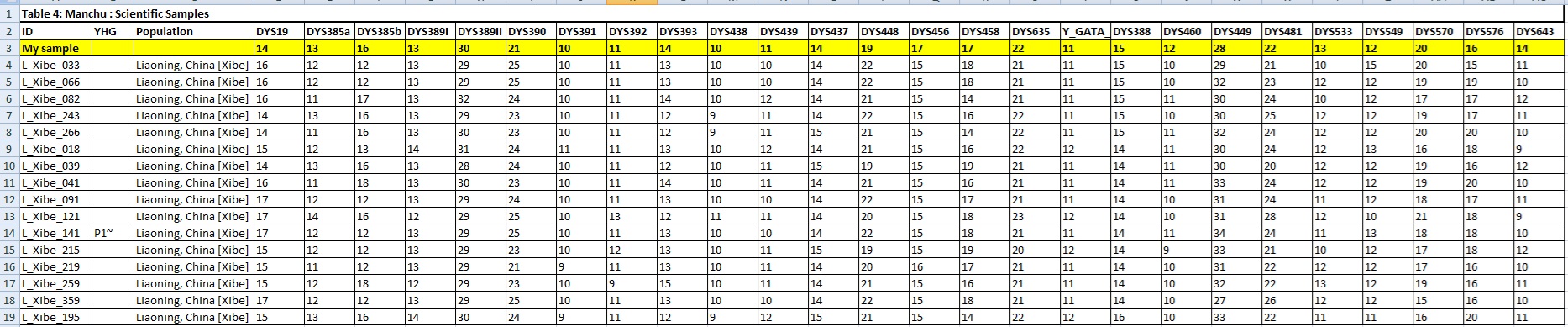
Y Lineage Confirmation: J*/J-Z7677/C-P53.1/O-Y3285
Y Lineage Origin: Ancient Caucasian/East Eurasian Steppe
-----------------------------------------------------------------------------------------------------------------
MTDNA Lineage Results and Confirmation: M37e-a1
MTDNA Origins: Ancient India / Middle East
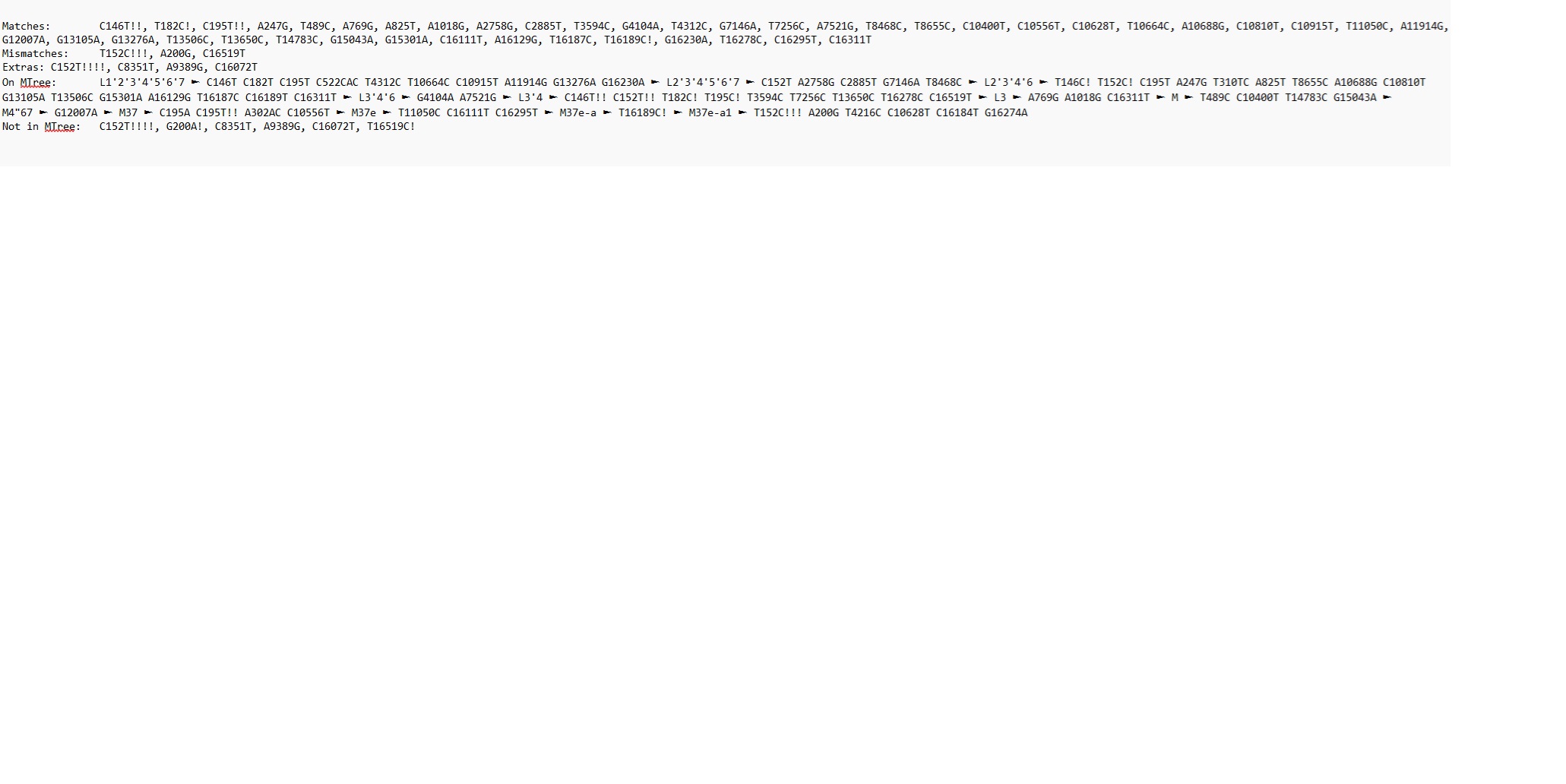
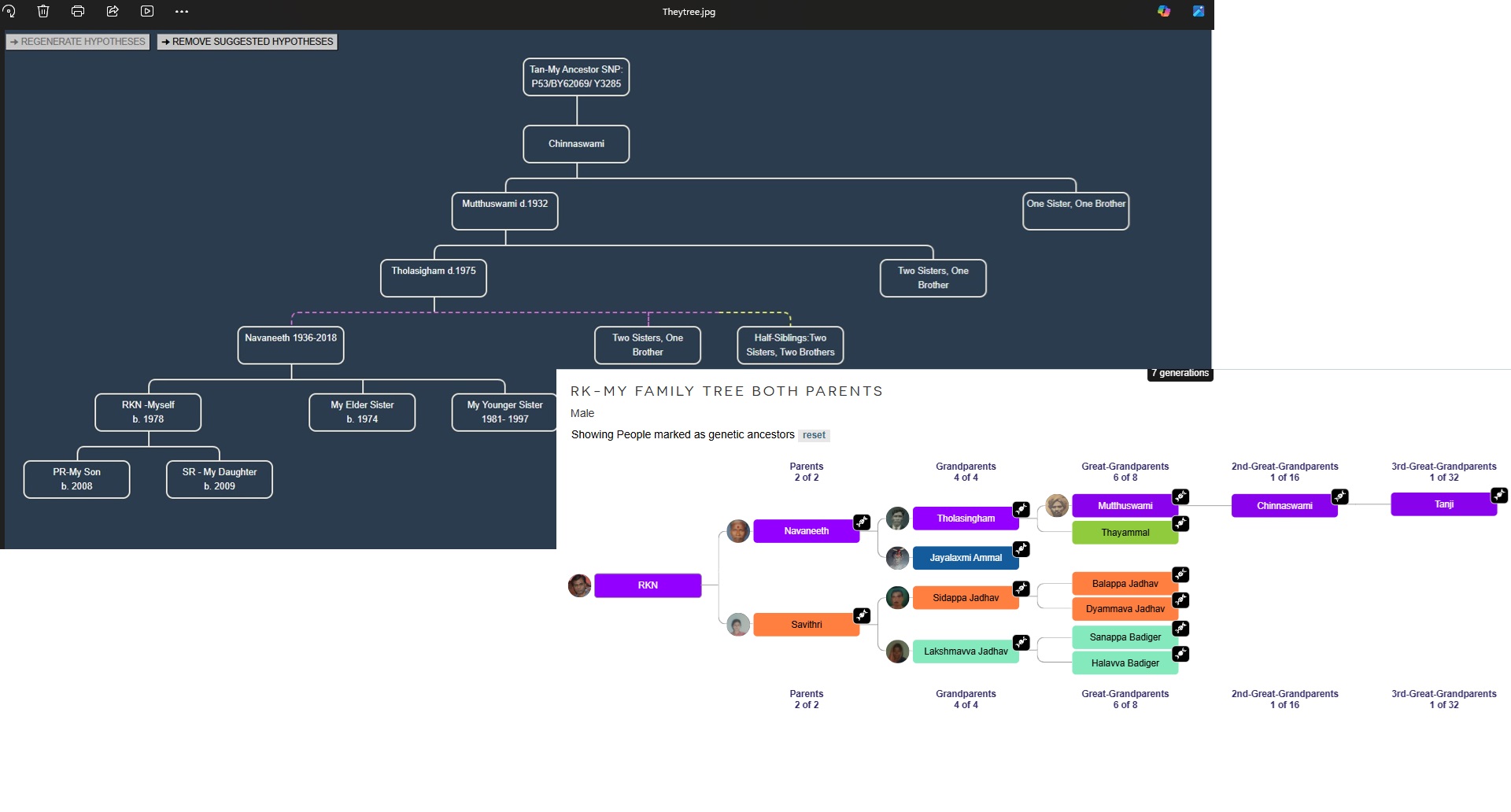
Current Location : India Y Lineage:Z13888/Y6718/FGC6800/BY62069/Z7677/P53/FT99321/M9060/MZ127/FT390416/Y146761/Y3285:Old Era Buddhist ancestor MT Lineage: M37e-a1 : India
© 2020-2026 TheYtree.com

留言 (0)